| Description | Participants | Summaries | Products |
|---|
Archived NIMBioS Working Group
Spatial Cell Simulations
Topic: Developing methods and improving tools for spatially-realistic cell simulations
Meeting dates: December 1-3, 2015; March 21-23, 2016; October 19-21, 2016; March 27-29, 2017
Organizers:
Robert F. Murphy, Computational Biology, School of Computer Science, Carnegie Mellon Univ.
James R. Faeder, Computational and Systems Biology, Univ. of Pittsburgh
Objectives: Systems biology emphasizes the creation of mathematical or computational models of biological systems such as cells and tissues, as a means both to integrate all available information and to make predictions about unmeasured mechanisms or behaviors. The vast majority of these models focus on metabolic or transcriptional regulation and do not consider spatial organization. Spatially realistic models are needed to keep pace with the vast amount of data that is being generated by high-resolution and live-cell microscopy and that has uncovered many new kinds of spatial organization and complexity. While some tools for spatial modeling do exist, a number of major limitations need to be overcome. The proposed working group will address the critical issues in creating realistic mathematical/computational simulations of the inner workings and dynamics of eukaryotic cells, especially by accurately simulating changes in shape and organization over time. The issues to be addressed include methods for simulation that can consider dynamic cell and organelle shapes and positions (movable boundary conditions) and methods for learning joint probability distributions for thousands of cellular components. The series of working group meetings will develop new approaches and implement them in software. Activities will likely also include development of proposals for funding in this area and development of training materials for biomedical researchers.

Meeting Summaries
| Mtg # | Dates | Agenda | Summary | Photo | Evaluation |
|---|---|---|---|---|---|
| 1 | Dec 1-3, 2015 | Link | Report | ||
| 2 | Mar 21-23, 2016 | Link | Link | ||
| 3 | Oct 19-21, 2016 | Link | Link | ||
| 4 | Mar 27-29, 2017 | Link | Link |
Meeting 2 Summary. The main activities of the Working Group's second meeting were 1) to identify a wish list for capabilities that the group would like to see general purpose spatial modeling tools address in the future and 2) to define a set of test problems relevant to spatial modeling of cellular and subcellular processes that can be used as benchmarks to compare various different packages and methods for simulation. This list includes very simple problems that should be solvable by any simulation program as well as complex problems that can be solved (at present) by only a few. Many of the problems can be formulated in either deterministic or stochastic modes to enable direct comparison of the consequences of this choice for simulation output. The various problems explore particular challenges in biological simulation including accurate rendering of complex temporal dynamics, spatial symmetry-breaking, interactions between combinations of 3D (soluble) and 2D (membrane-associated) species, and coupling between molecular motion and force generation. This list is not meant to be exhaustive, but rather to permit a manageably-sized survey of the kinds of problems that different simulation programs may be challenged to solve within a larger biological study. By framing the problems explicitly, the group aims to facilitate a direct comparison among different simulation packages both with respect to ease of encoding these particular scenarios and the speed and accuracy of execution. The group has set up a website at https://spatialmodel.wordpress.com/ that contains detailed descriptions of these test problems along with the wish list for spatial modeling capabilities. The group plans to further refine the test problems and to develop and post specific implementations for various publicly-available modeling tools for the next working group meeting. By framing the problems explicitly, the group aims to facilitate a direct comparison among different simulation packages both with respect to ease of encoding these particular scenarios and the speed and accuracy of execution.
Meeting 3 Summary. The group initially reviewed progress since the last meeting, which included implementation and testing of benchmark problems. Maggie Johnson presented simulation results for six different benchmark problems varying from simple binding to the establishment of spatial waves of concentration in the minCDE bacterial division system. Bob Murphy introduced his new cell geometry repository, and the group discussed how different instantiations of cell geometry can be used to investigate the accuracy of different tools and the sensitivity of the model to spatial variations. The group went on to discuss additional benchmark problems that would be useful for further testing the accuracy, flexibility, and capabilities of different spatial software tools. The group outlined a report that will discuss the current status of spatial cell modeling, the functionality of various tools, the challenges and the outlook for future progress. It will incorporate the results on benchmark problems to provide quantitative data for understanding the strengths and limitations of specific tools. The benchmark results are being shared and updated through github, and a public web site is being used to disseminate descriptions of the challenge problems and summaries of the benchmark results. Work is ongoing to provide the benchmark models in a variety of formats that will make the models simulatable in a wide range of available software frameworks.
Meeting 4 Summary. The major focus of this meeting was to begin drafting a report that will present the major findings and recommendations of the Spatial Cell Simulations Working Group. The group discussed how key several scientific challenges in cell biology highlight the need for further development of modeling and simulation tools. The group has developed a set of benchmark problems that can be simulated using current tools that will serve to highlight the capabilities of current tools and to compare their accuracy. The group plans to generate results for five different computational methods: ordinary differential equations (ODEs), stochastic simulation algorithm (SSA), partial differential equations (PDE), reaction-diffusion master equation (RDME), and single-particle simulation (SPS). Preliminary results are already available at the group's website, covering four of the five methods and five of the benchmark problems. The group also worked to identify a more challenging set of biological questions that reveal the limits of available simulation technology. The group discussed major issues arising from these challenges, which include lack of methods to address required biophysics, usability of the modeling software, scalability of simulation approaches, validation of simulation accuracy, and comparison of spatial data obtained from simulations and/or experiments. The group also discussed how the spatial modeling tools they develop could be further integrated both through continued development of standards and sharing of designs, algorithms, and code.
 |
| Meeting 1 participants (L to R): Carlos Lopez, Martin Meier-Schellersheim, Linda Petzold, Robert Murphy, Adelinde Uhrmacher, Richard Schugart, Julie Theriot, Ion I. Moraru, Margaret Johnson, James Faeder. |
 |
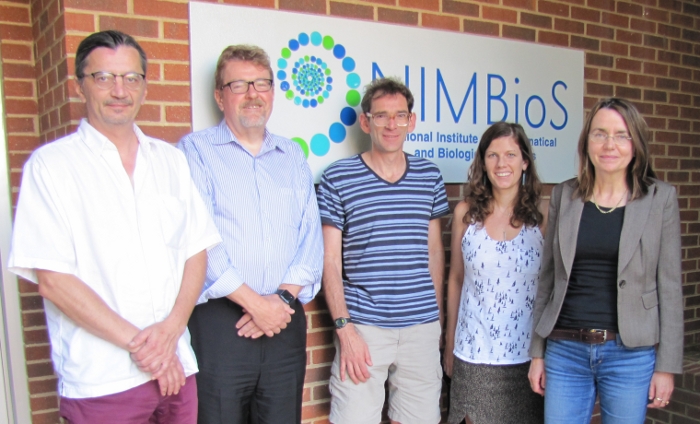
| Meeting 2 participants (L to R): Martin Meier-Schellersheim, James Faeder, Margaret Johnson, Alex Mogilner, Julie Theriot, Ion I. Moraru, Adelinde Uhrmacher. Not pictured: Igor Jouline, Richard Schugart. |
 |
| Meeting 3 participants (L to R): Ion I. Moraru, M, James Faeder, Margaret Johnson, Adelinde Uhrmacher |
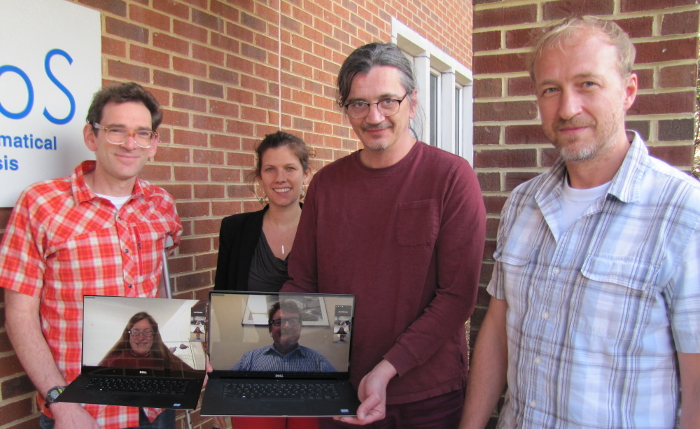 |
| Meeting 4 participants (L to R): James Faeder, Margaret Johnson, Ion Moraru, Martin Meier-Schellersheim, Julie Theriot (virtual), Robert Murphy (virtual) |
NIMBioS Working Groups are chosen to focus on major scientific questions at the interface between biology and mathematics. NIMBioS is particularly interested in questions that integrate diverse fields, require synthesis at multiple scales, and/or make use of or require development of new mathematical/computational approaches. NIMBioS Working Groups are relatively small (up to 10 participants), focus on a well-defined topic, and have well-defined goals and metrics of success. Working Groups will meet up to 3 times over a two-year period, with each meeting lasting up to 2.5 days.
A goal of NIMBioS is to enhance the cadre of researchers capable of interdisciplinary efforts across mathematics and biology. As part of this goal, NIMBioS is committed to promoting diversity in all its activities. Diversity is considered in all its aspects, social and scientific, including gender, ethnicity, scientific field, career stage, geography and type of home institution. Questions regarding diversity issues should be directed to diversity@nimbios.org. You can read more about our Diversity Plan on our NIMBioS Policies web page. The NIMBioS building is fully handicapped accessible.
NIMBioS
1122 Volunteer Blvd., Suite 106
University of Tennessee
Knoxville,
TN 37996-3410
PH: (865) 974-9334
FAX: (865) 974-9461
Contact NIMBioS


