| Description | Participants | Summaries | Products | Feature |
|---|
Archived NIMBioS Working Group
Cross-Topology Registration
Topic: Darwinian morphometrics: Cross-topology registration of shape
Organizers:
Patrick A. Carter (School of Biological Sciences, Washington State Univ.)
Richard Gomulkiewicz (Dept. of Mathematics and School of Biological Sciences, Washington State Univ.)
David Houle (Dept. of Biological Science, Florida State Univ.)
J. Steven Marron (Dept. of Statistics and Operations Research, Univ. of North Carolina, Chapel Hill)
Meeting dates: Jan. 10-12, 2010; Jan. 8-10, 2011; Jun. 13-15, 2012; May 1-3, 2013
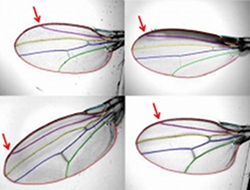
Project summary: Many complex traits of central importance in biology are defined by mathematical functions, and the variation, selection and evolution of these "function-valued traits" are of increasing interest. One fundamental challenge to understanding the evolution of function-valued traits is properly registering (or aligning) them while accounting for variation among individuals or taxa. This challenge is especially acute when considering morphological shapes, such as fly wings, but also occurs for other types of traits, such as ontogenies and reaction norms. Our view is that the problem of registration is not separate from analysis of the functions themselves, but rather is an essential part of the analysis. However, existing methods used by evolutionary biologists are not explicitly based on hypotheses about the nature of variation, but rather on convenient geometric properties that allow a general approach to be developed. We propose developing much deeper understanding of the actual biological processes that underlie differences in form, by the novel approach of integrating biological hypotheses directly into the geometric operation of registration and the resulting statistical analysis. Our proposed working group activities will synthesize the development of appropriate hypotheses with methods of registration not previously considered by evolutionary biologists, such as point distribution models, voxel based space warps, and medial models, to produce logical and systematic methods of analysis. We anticipate that our work will lead to multiple data analysis and theory papers, at least one summary paper describing the problem and its solutions, and computer program modules for general distribution and use. Four 3-day meetings are planned over the next 2 years.

Meeting Summaries
| Mtg # | Dates | Agenda | Summary | Photo | Evaluation |
|---|---|---|---|---|---|
| 1 | Jan 10-12, 2010 | Link | Link | Report | |
| 2 | Jan 8-10, 2011 | Link | Report | ||
| 3 | Jun 13-15, 2012 | ||||
| 4 | May 1-3, 2013 | Link |
Meeting 1 Summary. During the first meeting, empiricists described systems with which they work and specific and general statistical challenges for which they need solutions, while the statisticians described the types of problems central to their research and their general approach to solving those problems. Breakout groups were formed to focus on solving specific problems. Three major areas discussed included using Bayesian approaches to improve estimates of genetic parameters of function-valued phenotypes, incorporating the genetic relationship of individuals into methods of aligning curves, and developing computational and bioinformatic infrastructure necessary for the study of variation in morphology at a genome-wide scale.
Meeting 2 Summary. Major topics covered during the second meeting of the Darwinian Morphometrics working group were registration of growth curves, registration of morphological shapes, precision estimates of quantitative genetic parameters using Bayesian approaches, estimation and mapping of the nearly null space, and development of a framework to address questions focused on phenotypic data based on variation throughout the whole genome. A manuscript for TREE was also outlined, and progress was made on developing several other manuscripts in preparation related to the above issues, including a paper to highlight three different statistical methods the group has developed to register traits that can be described as curves.
 |
| Meeting 1 participants (Back row, L to R): John Stinchcombe, Fang Yao, Benedikt Halgrimsson, Daniel Gervini, Mark Kirkpatrick, Joel Kingsolver, Steve Marron; (Front row, L to R): Eladio Marquez, Patrick Carter, Jay Beder, Karin Meyer, Washington Mio, Nancy Heckman, Richard Gomulkiewicz; (Kneeling, L to R): David Houle and Sarang Joshi |
 |
| Mtg. 4 participants (L to R): Richard Gomulkiewicz; Eladio Marquez; Daniel Gervini; Jay Beder; Patrick Carter; Karin Meyer; Steve Marron; David Houle; Washington Mio |
NIMBioS Working Groups are chosen to focus on major scientific questions at the interface between biology and mathematics. NIMBioS is particularly interested in questions that integrate diverse fields, require synthesis at multiple scales, and/or make use of or require development of new mathematical/computational approaches. NIMBioS Working Groups are relatively small (up to 10 participants), focus on a well-defined topic, and have well-defined goals and metrics of success. Working Groups will meet up to 3 times over a two-year period, with each meeting lasting up to 2.5 days.
A goal of NIMBioS is to enhance the cadre of researchers capable of interdisciplinary efforts across mathematics and biology. As part of this goal, NIMBioS is committed to promoting diversity in all its activities. Diversity is considered in all its aspects, social and scientific, including gender, ethnicity, scientific field, career stage, geography and type of home institution. Questions regarding diversity issues should be directed to diversity@nimbios.org. You can read more about our Diversity Plan on our NIMBioS Policies web page. The NIMBioS building is fully handicapped accessible.
NIMBioS
1122 Volunteer Blvd., Suite 106
University of Tennessee
Knoxville,
TN 37996-3410
PH: (865) 974-9334
FAX: (865) 974-9461
Contact NIMBioS


