| Description | Participants | Summaries | Products | Report |
|---|
Archived NIMBioS Working Group
Agent-Based Models of Biological Systems
Topic: Agent-Based Models of Biological Systems: Pathways to control and optimization
Meeting dates: Apr. 26-28, 2011; Dec. 13-15, 2011; Nov. 27-29, 2012; Jan 21-24, 2014
Organizers:
• Gary An (Univ. of Chicago Pritzker School of Medicine)
• Reinhard Laubenbacher (Virginia Bioinformatics Institute)
• Suzanne Lenhart (Mathematics, Univ. of Tennessee, Knoxville)
• Jie Xiong (Mathematics, Univ. of Tennessee, Knoxville)
Objectives: This working group will explore mathematical and control/optimization frameworks and tools for agent-based/individual-based models (ABMs). The emphasis is on developing optimal control/optimization techniques that will work for a wide range of ABMs, addressing serious management and intervention problems. This group builds on the outcome of a workshop held at NIMBioS in December 2009. The workshop brought together a wide range of researchers, from domain experts in ecology, medicine, and epidemiology to mathematicians working in control theory and dynamical systems theory. Several clear themes emerged during the workshop, which will be addressed as part of the working group activities.
The group will examine several types of ABMs, based on characteristics of the ABM structure with respect to agent and world properties. This classification schema will form the basis of benchmark ABMs on which control/optimization techniques will be applied. Initially, two types of ABMs are defined, one with high numbers of agents with relatively simple and straightforward rules, and a second with smaller numbers of agents that have a high degree of rule complexity (manifest as simulated cognition and learning). Using agent-complexity as a metric, these two types of ABM establish the boundaries of a descriptive classification system for ABMs (recognizing that many if not most systems will be best represented by ABMs within the continuum). Aiming to develop robust and scalable approaches, two different application domains will be used as a preliminary test of the generalizability of the group's descriptive format and optimal control/optimization methods: the inflammatory response and ecosystems.
Having identified candidate ABMs that will serve as our test platforms, we will then apply a collection of mathematical representation and optimal control/optimization methods to each of these candidates and evaluate the relative success of each method as it pertains to each model ABM. The working group will focus initially on the development of investigatory teams centered around each method-application task:
- An algebraic representation of ABMs
- Approximation of ABMs with Optimal Control Techniques
- Stochastic Partial Differential Equations (SPDE) in Random Environments
- Optimization
Meeting Summaries
| Mtg # | Dates | Agenda | Summary | Photo | Evaluation |
|---|---|---|---|---|---|
| 1 | Apr 26-28, 2011 | Link | Link | Report | |
| 2 | Dec 19-21, 2011 | Link | |||
| 3 | Nov 27-29, 2012 | Link | |||
| 4 | Jan 21-24,2014 | Link | Link |
Meeting 1 Summary. The primary focus of the first meeting of this working group was on a careful definition of the problem area, which brings together many different possible mathematics and engineering approaches. These approaches are focused on the problem of developing effective methods for optimal control of biological systems, formalized as agent-based/ individual-based models (ABM/IBM). The biological systems under consideration are taken from biomedicine, in particular immunology, as well as ecology. The first part of the meeting was devoted to presentations by participants on their work in this problem area. The second part focused on the identification of several specific ABMs that can be used as test cases and as benchmark models for specific methods. Before the next meeting, working group participants will adapt their individual approaches to these case studies.
Meeting 2 Summary. The emphasis of the second meeting was on assessing the experiences of the various group members on applying mathematical formalisms to the reference ABMs selected at the first meeting, an ecological model and a biomedical/inflammation model, and using those experiences to further define the role of optimal control (OC) as applied to biology-based ABMs. Therefore the first part of the meeting was devoted to presentations by those groups that had worked on converting the reference ABMs to various mathematical methods, as well as operational strategies for assessing and comparing the resultant models. The first part of the meeting was also used to introduce new concepts concerning novel means of parsing ABMs to make then amenable to OC. The second part of the meeting included: developing a classification system for ABMs based on their incorporated properties to identify which aspects of an ABM could be suitably mapped to specific mathematical models, defining model reduction methods and the means to assess the fidelity of the reduced models, and the selection of reference ABMs based on these operational targets. Before the next meeting, working group participants will work on applying their various methods to the two reference models, with particular attention to defining suitable control functionals.
Meeting 3 Summary. This meeting focused on discussing ongoing projects, creating the outline for a position paper on ABMs, and planning for future work. The group has adapted the Netlogo Sugarscape model to include individuals who sense the landscape and move along a resource gradient and are taxed according to their "sugar" wealth. The group has made progress on building a difference equation model to approximate the Sugarscape ABM. Pareto optimization was used to rank "tax" control strategies. For the Sugarscape ABM with a more complex landscape, progress has been made on approximation using nonlinear difference equations and on optimization using genetic algorithms. For the Rabbits/Grass ABM model, a discrete time model counting the number of rabbits with the various energy levels was constructed to approximate the ABM, and control strategies were ranked with Pareto optimization. We have work in progress on partial differential equation approximation of those two ABMs and corresponding optimal control formulations.
Meeting 4 Summary. The fourth meeting comprised discussions on a number of topics, ranging from the polishing of certain dynamical systems approximations to promising new approaches to models for ABM control to organization and refinement of publication plans of attack. Two partial differential equation control models were finalized in this meeting, one for the Sugarscape ABM, using taxation as a control variable, and another for the rabbits and grass ecological model with rabbit harvesting controls. New approximation approaches included an empirical systems identification technique, which was successfully applied to the rabbits and grass ABM, and a controlled stochastic PDE system for the rabbits-grass interaction system. Publication plans include a review paper that examines current (mostly ad hoc) mathematical and computational methods to control of ABMs and proposes more unified approaches based on our Working Group’s research, as well as a number of research papers related to the results discussed in the meeting.
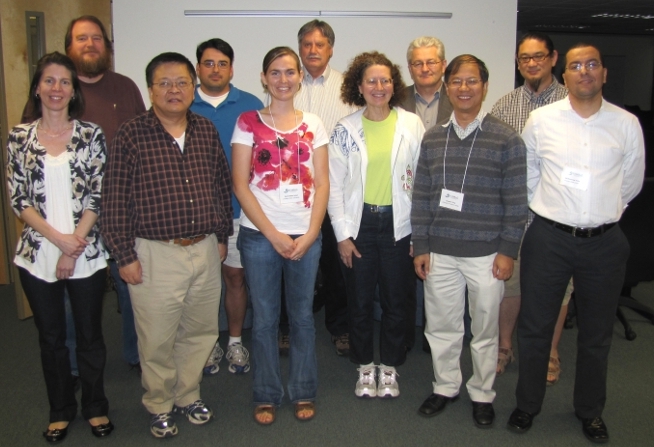 |
| Meeting 1 participants (Back row, L to R): Ben Fitzpatrick, René Salinas, Michael Bevers, Reinhard Laubenbacher, Gary An; (Front, L to R): Paula Federico, Jie Xiong, Rachel Miller Neilan, Suzanne Lenhart, Jiongmin Yong, Abdessamad Tridane |
 |
| Meeting 4 participants (L to R): Paula Federico, Suzanne Lenhart, Rene Salinas, Scott Christley, Matt Oremland, Andrew Kanarek (via Skype), Rachel Miller Neilan, Reinhard Laubenbacher, Jie Xiong, Ben Fitzpatrick, David Gurarie. Not pictured: Gary An. |
NIMBioS Working Groups are chosen to focus on major scientific questions at the interface between biology and mathematics. NIMBioS is particularly interested in questions that integrate diverse fields, require synthesis at multiple scales, and/or make use of or require development of new mathematical/computational approaches. NIMBioS Working Groups are relatively small (up to 10 participants), focus on a well-defined topic, and have well-defined goals and metrics of success. Working Groups will meet up to 3 times over a two-year period, with each meeting lasting up to 2.5 days.
A goal of NIMBioS is to enhance the cadre of researchers capable of interdisciplinary efforts across mathematics and biology. As part of this goal, NIMBioS is committed to promoting diversity in all its activities. Diversity is considered in all its aspects, social and scientific, including gender, ethnicity, scientific field, career stage, geography and type of home institution. Questions regarding diversity issues should be directed to diversity@nimbios.org. You can read more about our Diversity Plan on our NIMBioS Policies web page. The NIMBioS building is fully handicapped accessible.
NIMBioS
1122 Volunteer Blvd., Suite 106
University of Tennessee
Knoxville,
TN 37996-3410
PH: (865) 974-9334
FAX: (865) 974-9461
Contact NIMBioS


