2016 SRE Project
Decoding Allostery by Mathematical Analysis of Molecular Dynamics Simulations
Mentors:
Dr. Quentin Johnson, NIMBioS Postdoctoral Fellow
Dr. Tongye Shen, Biochemistry & Cellular and Molecular Biology (BCMB), UTK
Participants: Joshua Darville (Fisk Univ.), Elman Gonzales (East Tennessee State Univ.), and Jan Siess (Rutgers Univ.)
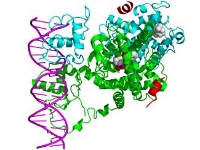 Allostery is a fundamental regulatory process, by which a biomolecule (or molecular complex) transmits a signal from one location to another distant site by a complex and seemly invisible molecular interaction network. This cooperative interaction (molecular cross-talk) has been found to be integral in many biological functions such as oxygen binding, enzyme regulation, and immune response. We will utilize a newly developed computational tool for the detection of allostery to investigate a nuclear receptor complex, a family of allosteric proteins that are key in transcriptional regulation. This project will involve using state of the art super computers to perform molecular dynamics simulations and analyze simulation trajectories using novel data reduction techniques. The goal of the project is to further our understanding of allostery in general and in advanced situations (such as negative allostery and promiscuous regulation); ultimately this work will lead to the future design of biomolecular switches.
Allostery is a fundamental regulatory process, by which a biomolecule (or molecular complex) transmits a signal from one location to another distant site by a complex and seemly invisible molecular interaction network. This cooperative interaction (molecular cross-talk) has been found to be integral in many biological functions such as oxygen binding, enzyme regulation, and immune response. We will utilize a newly developed computational tool for the detection of allostery to investigate a nuclear receptor complex, a family of allosteric proteins that are key in transcriptional regulation. This project will involve using state of the art super computers to perform molecular dynamics simulations and analyze simulation trajectories using novel data reduction techniques. The goal of the project is to further our understanding of allostery in general and in advanced situations (such as negative allostery and promiscuous regulation); ultimately this work will lead to the future design of biomolecular switches.

|
| Project group (from L): Joshua Darville, Elman Gonzales, Jan Siess |
Return to SRE 2016.
NIMBioS
1122 Volunteer Blvd., Suite 106
University of Tennessee
Knoxville,
TN 37996-3410
PH: (865) 974-9334
FAX: (865) 974-9461
Contact NIMBioS


