Explorations Through Time
Chai charts the course of evolutionary history
February 29, 2012
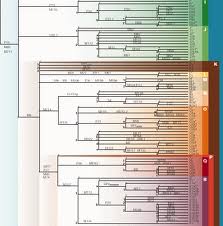
NIMBioS postdoctoral fellow Juanjuan "J.J." Chai is conducting research in phylogenetics, which studies evolutionary relatedness among groups of organisms through molecular sequencing data and morphological data. Biologists estimate that there are about 5 to 100 million species of organisms living on Earth today, and their genealogical relationships can be represented by evolutionary trees. Chai is working to solve problems related to the methods used in phylogenetics.
One of the methods used in the study of phylogenetics is maximum parsimony. Under parsimony, phylogenies are reconstructed from character sequences, such as DNA or morphological sequences, at the species level. The most parsimonious tree is the one that requires the fewest evolutionary changes for all sequences to derive from a common ancestor.
However, there are known problems in estimating phylogeny by using maximum parsimony, one being that the method can lead to different phylogenetic trees. And Chai is working to solve problems associated with maximum parsimony, as well as problems associated with other methods in phylogenetics.
Chai also builds mathematical models that try to explain biological mechanisms or phenomena. "With good models, we can do predictions of the behavior of processes, or pinpoint critical regions," she says. She is now working on a model to help understand how proteins evolve and incorporating results from population genetics.
Originally from Anhui, China, Chai has a Ph.D. in mathematics from Indiana University, Bloomington.
For more information about postdoctoral fellowships and other research and educational opportunities at NIMBioS, visit our website at http://www.nimbios.org.
#
The National Institute for Mathematical and Biological Synthesis (NIMBioS) brings together researchers from around the world to collaborate across disciplinary boundaries to investigate solutions to basic and applied problems in the life sciences. NIMBioS is supported by the National Science Foundation, the U.S. Department of Homeland Security, and the U.S. Department of Agriculture with additional support from The University of Tennessee, Knoxville.
NIMBioS
1122 Volunteer Blvd., Suite 106
University of Tennessee
Knoxville,
TN 37996-3410
PH: (865) 974-9334
FAX: (865) 974-9461
Contact NIMBioS


